Why does solid ice float on liquid water? Why are some batteries rechargeable and others not? How can plants turn their leaves to the sun? The remarkable behaviour of materials somehow emerges from their smallest constituents, the molecules, atoms, and electrons. But how this comes about is often shrouded in mystery and difficult to probe with experiments. Computer simulations and quantum chemical models are powerful tools to visualize the motions of the smallest particles, provide an interpretation to the experimental observations, and improve chemical and physical theories. Fundamental understanding of the dynamics of the microscopic details and how they yield the macroscopic properties is essential, for example to design better electro-catalysts to make sustainable fuels or capture CO2, to improve the performance of biodegradable plastics, and to develop specific drugs against diseases, to name but a few fields of application.
We use molecular simulations to answer fundamental questions on reaction mechanisms, properties of materials, and emergent behaviour in various (bio-) systems. We employ accurate quantum chemistry (DFT) based molecular dynamics simulations to study electron transfer and chemical reactivity, but also atomistic and coarse-grained forcefield simulations to investigate liquids, biomolecules, and soft materials on a larger scale. Recent research topics include (1) understanding the superior activity of hydrogenase enzymes compared to biomimetic catalysts for hydrogen fuel production, and subsequent development of catalyst design rules, (2) improving models for electro-catalytic water splitting, (3) understanding solvent behaviour in electron and proton transfer reactions, and (4) unravelling the photo-activation mechanism of light-sensing proteins.
Our arsenal of modelling tools includes a range of powerful algorithms and software that have been developed in the group throughout the years. Examples are PathCV and PathMap techniques to determine reaction mechanisms and compute reaction free energy landscapes, Hybrid-MD for adaptive resolution simulations, the Proton-Tracker-CV to model proton conductivity and determine acidity constants, and the FABULOUS method for finding reaction coordinates. Since recent years, we take advantage of machine learning developments that we adapt and exploit for data analysis (e.g. in FABULOUS) and surrogate modelling.
Research interests
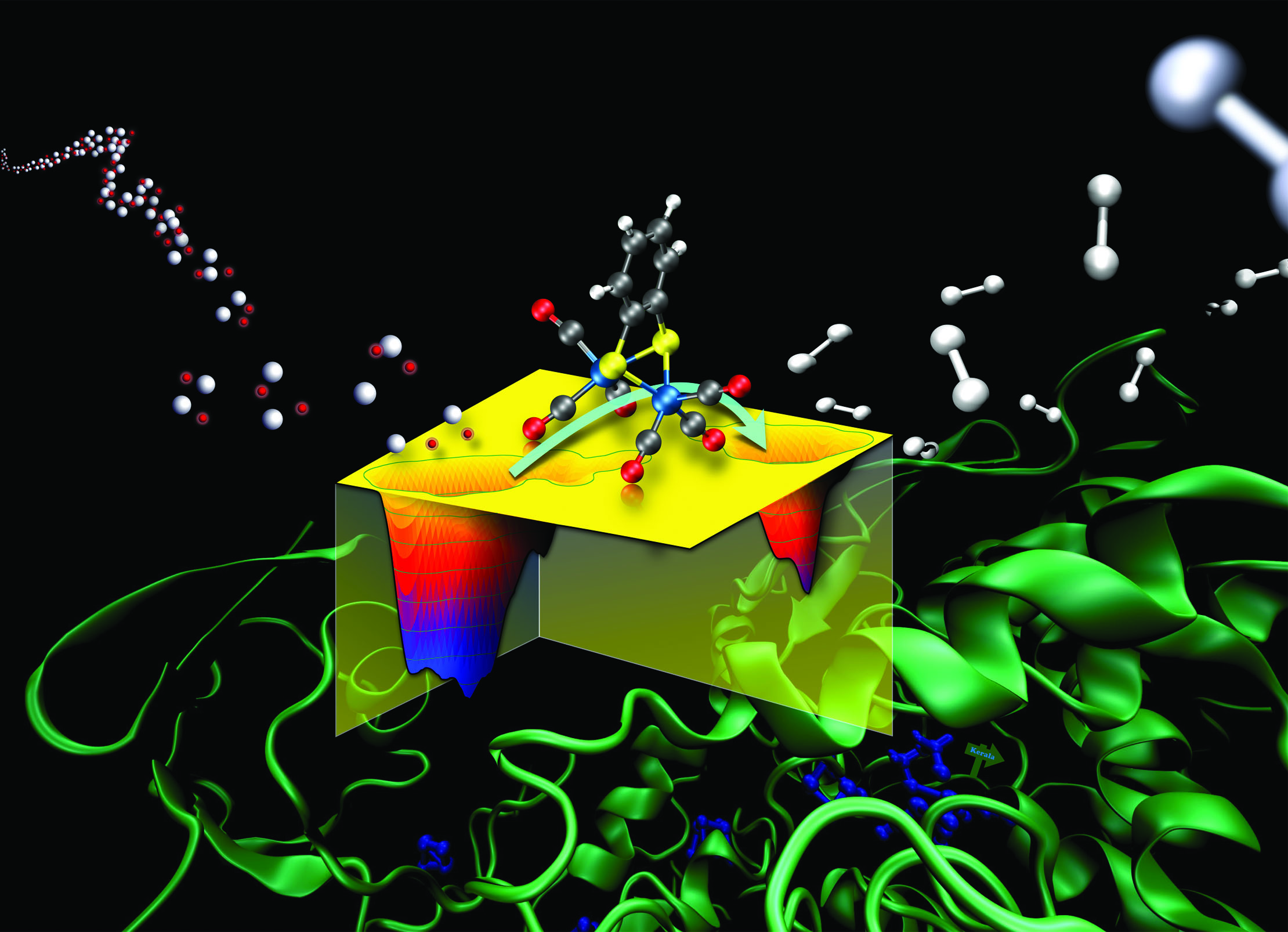
Bio-inspired catalysts for sustainable fuels and materials
Chemists have a central role in making our society sustainable and building a circular economy. Producing sustainable fuels, storing wind and solar energy, capturing CO2, and fixing nitrogen are all chemical processes that require better (electro-) catalysts. We use molecular simulations to learn design rules for new catalysts from natures most efficient enzymes.
Further reading:
-
Unraveling the mechanism of biomimetic hydrogen fuel production – A first principles molecular dynamics study.
Phys. Chem. Chem. Phys. 22, 10447 – 10454 (2020) -
A DFT study of structure and electrochemical properties of diiron-hydrogenase models with benzenedithiolato and benzenediselenato ligands.
New J. Chem. 44, 932 – 941 (2020)
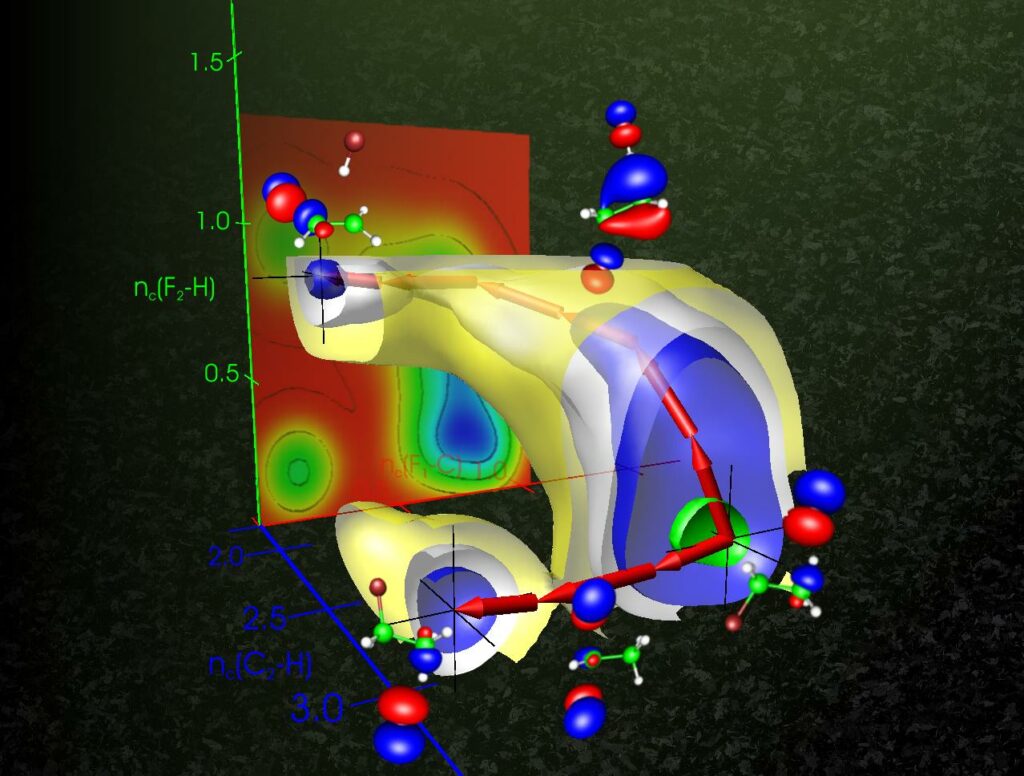
Path-metadynamics and PathMaps for free energy landscape exploration
We develop and apply enhanced sampling algorithms and software to simulate activated molecular processes that take place on long time scales, so-called rare events. Path-metadynamics, for example, is a technique to compute the reaction pathway and the free energy profile of chemical reactions, conformational transitions in biomolecules, etc. We develop machine learning algorithms to analyse complex transitions.
Further reading:
- Discovering collective variables of molecular transitions via genetic algorithms and neural networks. J. Chem. Theory Comput. 17, 2294 (2021)
- Advances in enhanced sampling along adaptive paths of collective variables. J. Chem. Phys. 149, 072320 (2018)
- Hydrogen activation by frustrated Lewis pairs revisited by metadynamics simulations. J. Phys. Chem. C 121, 2046 (2017)
- Metadynamics as a tool for exploring the free energy landscape of chemical reactions. Acc. Chem. Res. 39 , 73 (2006)
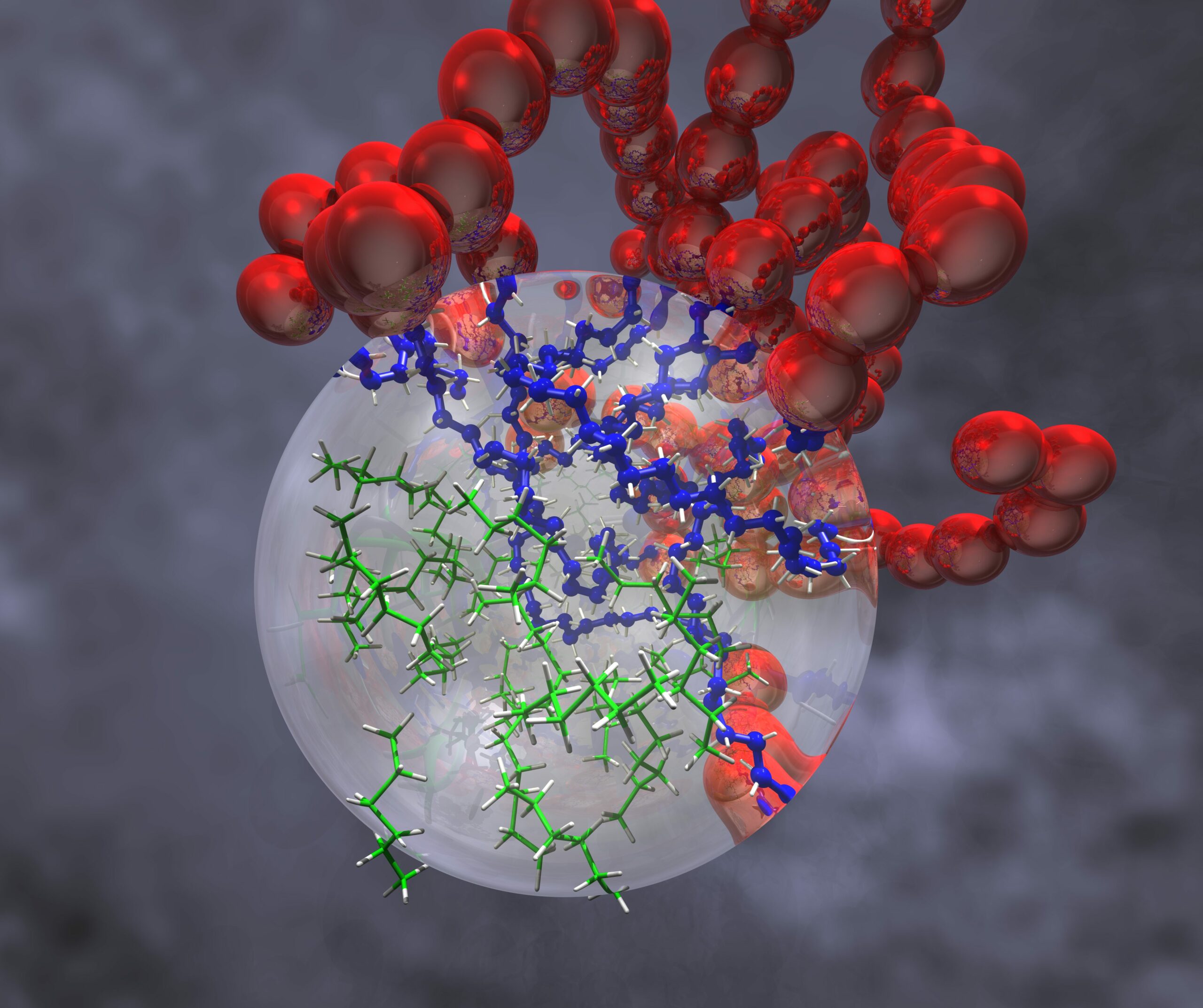
Multiscale modeling of soft advanced materials
Soft materials, such as biopolymers, coatings, polyelectrolytes, and fibres, display fascinating behavior at micro, meso, and macroscopic time and length scales. Polymers that adapt to external stimuli, known as smart materials, can be used in sensors or actuators. We use atomistic and coarse-grained simulations to model the behaviour of polymers and polymer surfaces and study how polymer systems can be tailored with tunable functionalities, such as durability, anti-fouling, self-healing, or biodegradability. We develop hybrid atomistic/coarse-grain simulation techniques to model large systems in chemical detail.
Further reading:
- Double helical conformation and extreme rigidity in a rodlike polyelectrolyte.
Nature Communications 10, 801 (2019) - On the “tertiary Structure” of poly-Carbenes; self-assembly of sp3-carbon based polymers into liquid-crystalline aggregates.
Chem. Eur. J. 19, 11577 (2013) - Recent progress in multiscale molecular dynamics simulation of soft matter.
Phys. Chem. Chem. Phys. 12, 12401 (2010) - Adaptive multiscale molecular dynamics of macromolecular fluids.
Phys. Rev. Lett. 105, 237802 (2010)
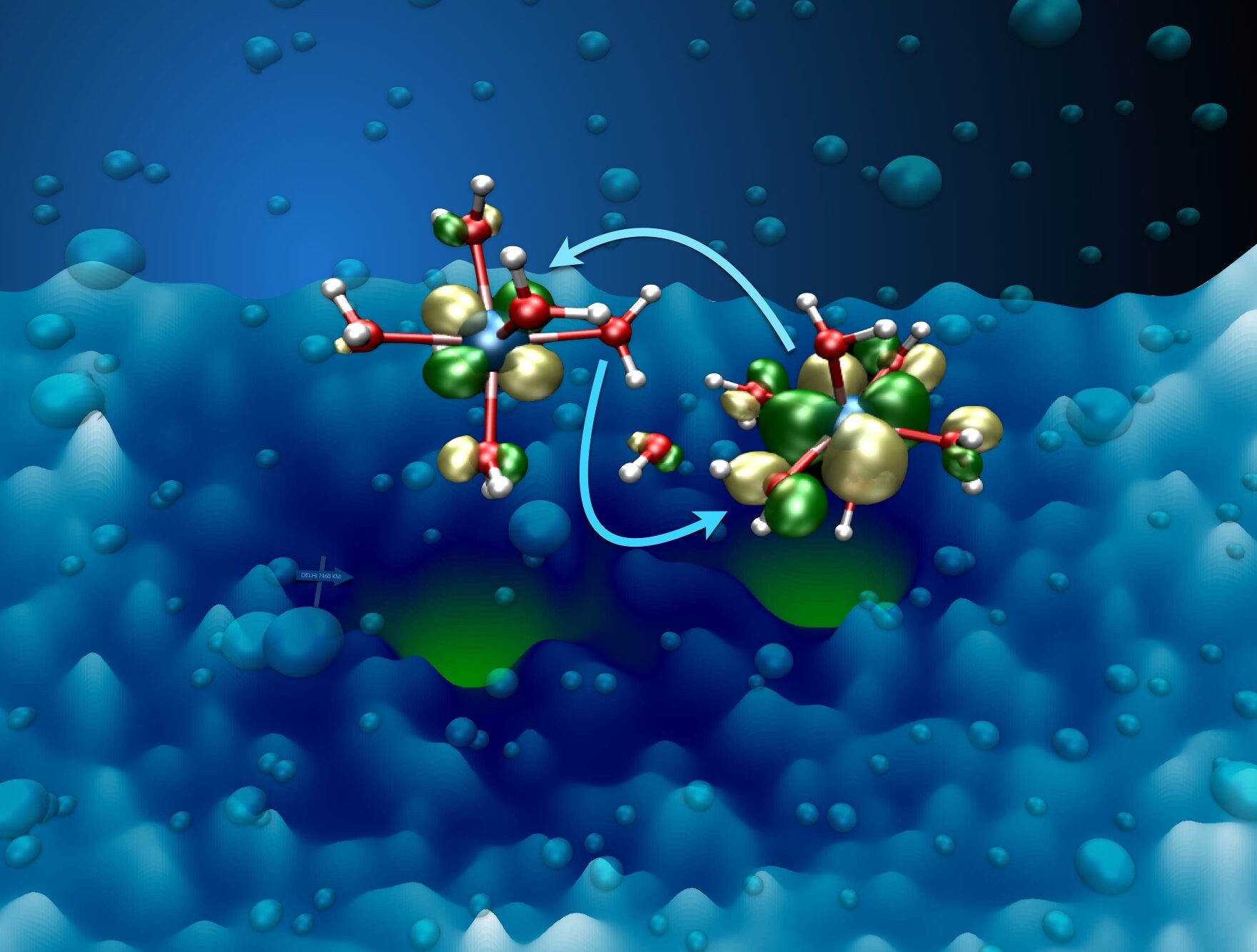
Electron and proton transfer in solution
Protons and electrons are very light particles that can move with a high velocity, but in practice their transfer kinetics is strongly affected by the medium, which can e.g. be a solvent, a protein assembly, or an electrode-electrolyte interface. The conductivity in a solvent is governed by the solvent fluctuations. Confinement of the solvent in one or more dimensions affects the transfer mechanism. The solvent reorganization free energy is an important part of both the redox potential and the acidity constants of acceptor solutes.
Further reading
- Reactive trajectories of the Ru2+/3+ self-exchange reaction and the connection to Marcus’ theory.
Faraday Discuss. 195, 291 – 310 (2016) - Microscopic Picture of the Solvent Reorganization During Electron Transfer to Flavin in Water.
J. Phys. Chem. B 123, 9751 – 9761 (2019) - Acidity constants of lumiflavin from first principles molecular dynamics simulations.
Phys. Chem. Chem. Phys. 16 , 18993 (2014)
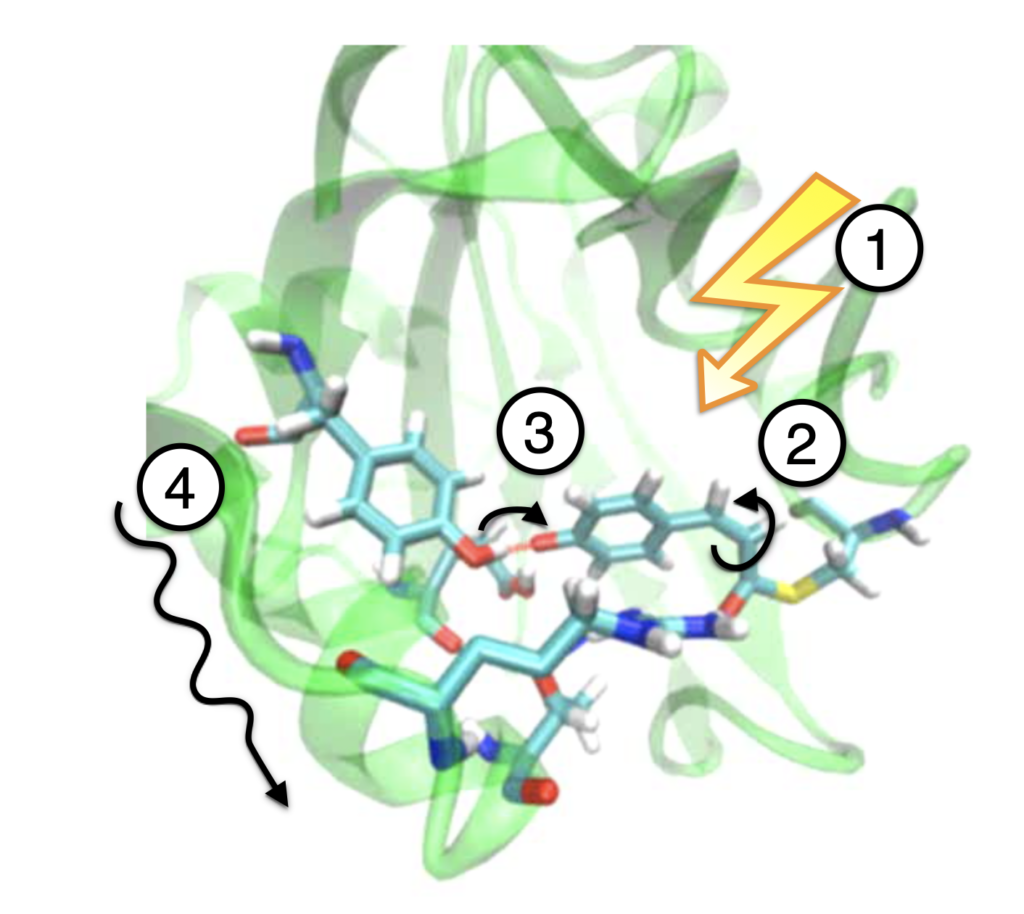
Signal transduction in photoactive proteins
All organisms have sensor proteins that allow them to respond to changes in their environment. These proteins can sense for example pressure, temperature, or chemical compounds and they are grouped in protein families that share similar structures (or folds) and can bind to similar output domains. Light sensing proteins are particularly interesting for studying their mechanisms, because they can be triggered with a flash of laser light and then probed for internal changes. Using enhanced DFT (QM), forcefield (MM), and mixed QM/MM simulations, we see how such photoactive proteins undergo chemical and conformational changes when exposed to light. Our simulations help with interpreting spectroscopic experiments and with understanding for example why some protein family members have much longer signalling times than others.
Further reading
- Frontiers in Multiscale Modelling of Photoreceptor Proteins.
Photochem. Photobiol. 97, 243 – 269 (2020)

The extraordinary behaviour of water and aqueous systems
-
On the origin of the extremely different solubilities of polyethers in water.
Nature Communications 10, 2893 (2019) -
A liquid-liquid transition in supercooled aqueous solution related to the HDA-LDA transition.
Science 359 , 1127 (2018) -
On the slowdown mechanism of water dynamics around small amphiphiles.
Phys. Chem. Chem. Phys. 17, 24968 (2015) -
Local orientational ordering in liquids revealed by resonant vibrational energy transfer.
Phys. Rev. Lett. 113, 207801 (2014)
Publications
2025
96) Simultaneous sampling of multiple transition channels using adaptive paths of collective variables.
Alberto Pérez de Alba Ortíz and Bernd Ensing
J. Chem. Phys. 163, 144104 (2025)
Open-access preprint: arXiv 2112.04061v2 (2025)
95) Learned Free-Energy Functionals from Pair-Correlation Matching for Dynamical Density Functional Theory.
Karnik Ram, Jacobus Dijkman, René van Roij, Jan-Willem van de Meent, Bernd Ensing, Max Welling, Daniel Cremers
Phys. Rev. E 112, 045314 (2025)
Open-access preprint: arXiv 2505.09543 (2025)
94) Learning Neural Free-Energy Functionals with Pair-Correlation Matching.
Jacobus Dijkman, Marjolein Dijkstra, René van Roij, Max Welling, Jan-Willem van de Meent, and Bernd Ensing
Phys. Rev. Lett. 134, 056103 (2025)
Open-access preprint: arXiv 2403.15007 (2024)
2024
93) PLUMED Tutorials: a collaborative, community-driven learning ecosystem.
Gareth A. Tribello, Massimiliano Bonomi, Giovanni Bussi, et al.
J. Chem. Phys. 162, 092501 (2025)
Open-access preprint: arXiv 2412.03595 (2024)
92) Comparison of Optimization Algorithms for Automated Method Development of Gradient Profiles.
Gerben B. van Henten, Jim Boelrijk, Céline Kattenberg, Tijmen S. Bos, Bernd Ensing, Patrick Forré, Bob W.J. Pirok
J. Chromatogr. A, 1742 465626 (2025)
91) Redox Properties of Flavin in BLUF and LOV Photoreceptor Proteins from Hybrid QM/MM Molecular Dynamics Simulation.
Murat Kılıç and Bernd Ensing
J. Phys. Chem. B 128, 3069 – 3080 (2024)
Open-access preprint: ChemRxiv. Cambridge (2023)
90) Enhancing LC×LC separations through multi-task Bayesian optimization.
Jim Boelrijk, Stef R.A. Molenaar, Tijmen S. Bos, Tina A. Dahlseid, Bernd Ensing, Dwight R. Stoll, Patrick Forré, and Bob W.J. Pirok
J. Chromatogr. A 1726, 464941 (2024)
Open-access preprint: ChemRxiv. Cambridge 10.26434/chemrxiv-2024-5mmvj (2024)
2023
89) Stochastic Optimal Control for Collective Variable Free Sampling of Molecular Transition Paths.
Lars Holdijk, Yuanqi Du, Ferry Hooft, Priyank Jaini, Bernd Ensing, and Max Welling
Thirty-seventh Conference on Neural Information Processing Systems (NeurIPS) (2023)
Open-access preprint: arXiv: 2207.02149 (2022)
88) Predicting RP-LC retention indices of structurally unknown chemicals from mass spectrometry data.
Jim Boelrijk, Denice van Herwerden, Saer Samanipour, Bernd Ensing, and Patrick Forré
J. Cheminform. 15, 28 (2023)
Open-access preprint: ChemRxiv. Cambridge (2022)
87) Free energies of the Gln tautomerization and rotation mechanism of dark-state recovery in blue light-using flavin proteins.
Alberto Pérez de Alba Ortíz, Carme Rovira, and Bernd Ensing
bioRxiv: 10.1101/2023.08.11.551373 (2023)
86) Simulation of solid-state phase transition in DL-methionine.
Saba Ghasemlou, Bernd Ensing, and Herma M. Cuppen
CrystEngComm 25, 3618 – 3627 (2023)
85) Closed-loop automatic gradient design for liquid chromatography using Bayesian optimization.
Jim Boelrijk, Bernd Ensing, Patrick Forré, and Bob Pirok
Analytica Chimica Acta 1242, 340789 (2023)
Open-access preprint: ChemRxiv. Cambridge (2022)
2022
84) Multi-objective optimization via equivariant deep hypervolume approximation.
Jim Boelrijk, Bernd Ensing, and Patrick Forré
ICLR, The Eleventh International Conference on Learning Representations (2023)
Open-access preprint: arXiv: 2210.02177 (2022)
83) Chemometric strategies for fully automated interpretive method development in liquid chromatography.
Tijmen Bos, Jim Boelrijk, Stef Molenaar, Brian van ‘t Veer, Leon Niezen, Denice van Herwerden, Saer Samanipour, Dwight Stoll, Patrick Forré, Bernd Ensing, Govert Somsen, and Bob Pirok
Anal. Chem. 94 16060 – 16068 (2022)
Open-access preprint: ChemRxiv. Cambridge (2022)
82) Sequence dependence of transient Hoogsteen base pairing in DNA.
A. Pérez de Alba Ortíz, Jocelyne Vreede, and B. Ensing
PLOS Comp. Biol. 18, e1010113 (2022)
Open-access
81) Fast proton transport in FeFe hydrogenase via a flexible channel and a proton hole mechanism.
Rakesh Puthenkalathil and Bernd Ensing
J. Phys. Chem. B 126, 403 – 411 (2021)
Open-access
2021
80) Linear scaling relationships to predict pKa’s and reduction potentials for bio-inspired hydrogenase catalysis.
Rakesh Puthenkalathil and Bernd Ensing
Inorg. Chem. 61, 113 – 120 (2021)
79) Simultaneous sampling of multiple transition channels using adaptive paths of collective variables.
Alberto Pérez de Alba Ortíz and Bernd Ensing
arXiv [cond-mat.stat-mech] 2112.04061 (2021)
78) Bayesian Optimization of Comprehensive Two-dimensional Liquid Chromatography Separations.
Jim Boelrijk, Bob Pirok, Bernd Ensing, and Patrick Forré
J. Chromatogr. A 1659, 462628 (2021)
77) Computational study of electron transport in [FeFe] hydrogenase enzymes.
Rakesh Puthenkalathil and Bernd Ensing
ChemRxiv. Cambridge (2021)
76) Discovering collective variables of molecular transitions via genetic algorithms and neural networks.
Ferry Hooft, Alberto Pérez de Alba Ortíz, and Bernd Ensing
J. Chem. Theory Comput. 17, 2294 – 2306 (2021)
75) Strong reduction of the chain rigidity of hyaluronan by selective binding of Ca2+ ions.
G. Giubertoni, A. Pérez de Alba Ortíz, F. Bano, D. Green, P. DeAngelis, G.H. Koenderink, R. Richter, B. Ensing, and H.J. Bakker
Macromolecules 54, 1137 – 1146 (2021)
2020
74) Frontiers in Multiscale Modelling of Photoreceptor Proteins.
Maria-Andrea Mroginski et al.
Photochem. Photobiol. 97, 243 – 269 (2020)
73) Hydration interactions beyond the first solvation shell in aqueous phenolate solution.
Roberto Cota, Ambuj Tiwari, Bernd Ensing, Huib J. Bakker, and Sander Woutersen
Phys. Chem. Chem. Phys. 22 , 19940 – 19947 (2020)
72) Mechanistic Insight into the Hydrogen Activation by Frustrated Lewis Pairs.
Mojgan Heshmat, Lei Liu, and Bernd Ensing
Book chapter in Frustrated Lewis Pairs, Book series: Molecular Catalysis, J.C. Slootweg and A. Jupp. (eds), Springer International Publishing 2 (2021), 167 – 208
71) Optimizing the Energetics of FLP-Type H2 Activation by Modulating Electronic and Structural Properties of the Lewis Acids: A DFT Study.
Mojgan Heshmat and Bernd Ensing
J. Phys. Chem. A 124 , 6399 – 6410 (2020)
70) Peptide side-COOH groups have two distinct conformations under bio-relevant conditions.
Oleksandr Sofronov, Giulia Giubertoni, Alberto Pérez de Alba Ortíz, Bernd Ensing, Huib Bakker
J. Phys. Chem. Lett. 11 , 3466 – 3472 (2020)
69) Unraveling the mechanism of biomimetic hydrogen fuel production – A first principles molecular dynamics study.
Rakesh C. Puthenkalathil, Mihajlo Etinski, and Bernd Ensing
Phys. Chem. Chem. Phys. 22, 10447 – 10454 (2020)
68) A DFT study of structure and electrochemical properties of diiron-hydrogenase models with benzenedithiolato and benzenediselenato ligands.
Mihajlo Etinski, Ivana M. Stankovic, Rakesh C. Puthenkalathil, and Bernd Ensing
New J. Chem. 44, 932 – 941 (2020)
2019
67) Accurate calculation of zero point energy from molecular dynamics simulations of liquids and their mixtures.
Ambuj Tiwari, Coen Honingh, and Bernd Ensing
J. Chem. Phys. 151, 244124 (2019)
66) Microscopic Picture of the Solvent Reorganization During Electron Transfer to Flavin in Water.
Murat Kılıç and Bernd Ensing
J. Phys. Chem. B 123, 9751 – 9761 (2019)
65) The PLUMED consortium: A community effort to promote openness, transparency and reproducibility in molecular simulations.
The PLUMED consortium*
Nature Methods 16, 670 – 673 (2019)
64) On the origin of the extremely different solubilities of polyethers in water.
Bernd Ensing, Martijn Tros, Ambuj Tiwari, Johannes Hunger, Sergio Rosa Domingos, Christobal Perez, Gertien Smits, Mischa Bonn, Daniel Bonn, and Sander Woutersen
Nature Communications 10, 2893 (2019)
63) Cooperative Role of Water Molecules During the Initial Stage of Water-induced Zeolite Dealumination.
Katarina Stanciakova, Bernd Ensing, Florian Gӧltl, Rosa Bulo, and Bert Weckhuysen
ACS Catalysis 9, 5119 – 5135 (2019)
62) Double Helical Conformation and Extreme Rigidity in a Rodlike Polyelectrolyte.
Ying Wang, Yadong He, Zhou Yu, Jianwei Gao, Stephanie T. Brinck, Carla Slebodnick, Gregory B. Fahs, Curt J. Zanelotti, Maruti Hegde, Robert B. Moore, Bernd Ensing, Theo J. Dingemans, Rui Qiao, and Louis A. Madsen
Nature Communications 10, 801 (2019)
61) Reaction Mechanism of Hydrogen Activation by Frustrated Lewis Pairs.
Lei Liu, Binit Lukose, Pablo Jaque, Bernd Ensing
Green Energy & Environment 4, 20 – 28 (2019)
60) The Adaptive Path Collective Variable: A Versatile Biasing Approach to Compute the Average Transition Path and Free Energy of Molecular Transitions.
Alberto Pérez de Alba Ortíz, Jocelyne Vreede, Bernd Ensing
Book chapter in Biomolecular Simulations. Methods in Molecular Biology, Bonomi M., Camilloni C. (eds), Humana, New York, NY 2022 (2019), 255 – 290
2018
59) The Puzzle of the Intramolecular Hydrogen Bond of Dibenzoylmethane Resolved by Molecular Dynamics Simulations.
Mihajlo Etinski and Bernd Ensing
J. Phys. Chem. A 122, 5945 – 5954 (2018)
58) Advances in enhanced sampling along adaptive paths of collective variables.
Alberto Pérez de Alba Ortíz, Ambuj Tiwari, Rakesh C. Puthenkalathil, and Bernd Ensing
J. Chem. Phys. 149, 072320 (2018)
57) Mechanisms behind the enhancement of thermal properties of graphene nanofluids.
Maria del Rocio Rodríguez-Laguna, Alejandro Castro-Alvarez, Marinanna Sledzinska, Jeremie Maire, Francesca Costanzo, Bernd Ensing, Miguel Pruneda, Pablo Ordejón, Clivia M. Sotomayor Torres, Pedro Gómez-Romero and Emigdio Chávez-Ángel
Nanoscale 10, 15402 – 15409 (2018)
56) Impact of Ligand Flexibility and Solvent on the O-O bond formation step in a highly active Ru Water Oxidation Catalyst.
Nitish Govindarajan, Ambuj Tiwari, Bernd Ensing, and Evert Jan Meijer
Communication in Inorg. Chem. 57, 13063 – 13066 (2018)
55) A liquid-liquid transition in supercooled aqueous solution related to the HDA-LDA transition.
Sander Woutersen, Bernd Ensing, Michiel Hilbers, Zuofeng Zhao, and C. Austen Angell
Science 359, 1127-1131 (2018)
54) A free energy landscape of CO2 capture by frustrated Lewis pairs.
Lei Liu, Binit Lukose, and Bernd Ensing
ACS Catalysis 8, 3376-3381 (2018)
53) Designing effective solid catalysts for biomass conversion: Aerobic oxidation of ethyl lactate to ethyl pyruvate.
Wei Zhang, Bernd Ensing, Gadi Rothenberg and Raveendran N Shiju
Advance Article in Green Chemistry 20, 1866-1873 (2018)
52) Highly selective oxidation of ethyl lactate to ethyl pyruvate catalysed by mesoporous vanadia–titania.
Wei Zhang, Giada Innocenti, Paula Oulego, Vitaly Gitis, Hai-Hong Wu; Bernd Ensing, Fabrizio Cavani, Gadi Rothenberg, N. Raveendran Shiju
ACS Catalysis 8, 2365–2374 (2018)
2017
51) Hydrogen activation by frustrated Lewis pairs revisited by metadynamics simulations.
Lei Liu, Binit Lukose and Bernd Ensing
J. Phys. Chem. C 121, 2046 – 2051 (2017)
2016
50) Reactive trajectories of the Ru2+/3+ self-exchange reaction and the connection to Marcus’ theory.
Ambuj Tiwari and Bernd Ensing
Cover article in Faraday Discuss. 195, 291 – 310 (2016)
49) Acidity constant (pKa) calculation of large solvated dye molecules: evaluation of two advanced molecular dynamics methods.
Thierry De Meyer, Bernd Ensing, Sven M. J. Rogge, Karen De Clerck, Evert Jan Meijer and Veronique Van Speybroeck
ChemPhysChem 17, 3447 – 3459 (2016)
48) Energy barriers and mechanisms in solid-solid polymorphic transitions exhibiting cooperative motion.
Joost A. van den Ende, Bernd Ensing, and Herma M. Cuppen
CrystEngComm 18, 4420-4430 (2016)
47) Insight in the effect of water on the methanol-to-olefins conversion in H-SAPO-34 from molecular simulations and in-situ micro-spectroscopy.
Kristof De Wispelaere, Caterina S. Wondergem, Bernd Ensing, Karen Hemelsoet, Evert Jan Meijer, Bert M. Weckhuysen, Veronique Van Speybroeck, and Javier Ruiz-Martínez
ACS Catalysis 6, 1991-2002 (2016)
2015
46) On the slowdown mechanism of water dynamics around small amphiphiles.
Wagner H. Brandeburgo, Sietse T. van der Post, Evert Jan Meijer, and Bernd Ensing
Phys. Chem. Chem. Phys. 17, 24968 – 24977 (2015)
45) Complex reaction environments and competing reaction mechanisms in zeolite catalysis: insights from advanced molecular dynamics.
Kristof De Wispelaere, Bernd Ensing, An Ghysels, Evert Jan Meijer, and Veronique Van Speybroeck
Chem. Eur. J. 21, 9385 – 9396 (2015)
44) Do solid-to-solid polymorphic transitions in DL-norleucine proceed through nucleation?
Joost A. van den Ende, Mireille M. H. Smets, Daniël T. de Jong, Sander J. T. Brugman, Bernd Ensing, Paul T. Tinnemans, Hugo Meekes, and Herma M. Cuppen
Faraday Discuss. 179, 421 – 436 (2015)
43) Understanding DABCO nanorotor dynamics in isostructural metal-organic frameworks.
Nicholas Burtch, Ariana Torres-Knoop, Guo Shiou Foo, Johannes Leisen, Carsten Sievers, Bernd Ensing, David Dubbeldam, Krista Walton
J. Phys. Chem. Lett. 6, 812 – 816 (2015)
2014
42) Local orientational ordering in liquids revealed by resonant vibrational energy transfer.
Matthijs R. Panman, Daniel J. Shaw, Bernd Ensing, and Sander Woutersen
Phys. Rev. Lett. 113, (2014), 207801
41) Acidity constants of lumiflavin from first principles molecular dynamics simulations.
Murat Kılıç and Bernd Ensing
Phys. Chem. Chem. Phys. 16, (2014), 18993 – 19000
40) Interaction of H2 with a double walled armchair nanotube by first principles calculations.
Francesca Costanzo, Bernd Ensing, Roberto Scipioni, Francesco Ancilotto, and Pier Luigi Silvestrelli
J. Phys. Chem. C 118, 15816 – 15824 (2014)
39) Comment on Communication: Benzene dimer – The free energy landscape [J. Chem. Phys. 139, 201102 (2013)].
Ad van der Avoird, Rafal Podeszwa, Bernd Ensing and Krzysztof Szalewicz
J. Chem. Phys. 140, 227101 (2014)
2013
38) First and second one-electron reduction of lumiflavin in water – A first principles molecular dynamics study.
Murat Kılıç and Bernd Ensing
J. Chem. Theory Comput. 9, 3889 – 3899 (2013)
37) On the “tertiary Structure” of poly-Carbenes; self-assembly of sp3-carbon based polymers into liquid-crystalline aggregates.
Nicole M. G. Franssen, Bernd Ensing, Maruti Hegde, Theo Dingemans, Ben Norder, Stephen J. Picken, Gert O. R. Alberda van Ekenstein, Ernst van Eck, Johannes A. A. W. Elemans, Mark Vis, Joost N. H. Reek and Bas de Bruin
Cover article in Chem. Eur. J. 19, 11577 – 11589 (2013)
36) Hamiltonian adaptive hybrid atomistic/coarse-grain molecular dynamics.
Bernd Ensing, Alessandro Laio, and Steven O. Nielsen
Publication series of the John von Neumann Institute for Computing 46, 95 – 110 (2013)
2012
35) On the polarity of buckminsterfullerene with a water molecule inside.
Bernd Ensing, Francesca Costanzo, and Pier Luigi Silvestrelli
J. Phys. Chem. A 116, 12184 – 12188 (2012)
34) Path finding on high-dimensional free energy landscapes.
Grisell Díaz Leines and Bernd Ensing
Phys. Rev. Lett. 109, 020601 (2012)
2011
33) Quantitative assessment of force fields on both low-energy conformational basins and transition-state regions of the (φ, ψ) space.
Zhiwei Liu, Bernd Ensing, and Preston B. Moore
J. Chem. Theory. Comput. 7, 402 – 419 (2011)
32) A Reply to the Comment by M. Praprotnik et al.
Steven O. Nielsen, Preston B. Moore, Bernd Ensing
Phys. Rev. Lett. 107, 099802 (2011)
2010
31) Recent progress in multiscale molecular dynamics simulation of soft matter.
Steven O. Nielsen, Rosa E. Bulo, Preston B. Moore, Bernd Ensing
Invited Perspective and cover article Phys. Chem. Chem. Phys. 12, 12401 – 12412 (2010)
30) Adaptive multiscale molecular dynamics of macromolecular fluids.
Steven O. Nielsen, Preston B. Moore, and Bernd Ensing
Phys. Rev. Lett. 105, 237802 (2010)
29) Smooth capping of short-range repulsive forces in hybrid atomistic/coarse-grain molecular dynamics simulation.
Bernd Ensing
Proceedings of Multiscale Materials Modeling 5, 47 – 53 (2010)
28) Non-linear reaction coordinate analysis in the reweighted path ensemble.
Wolfgang Lechner, Jutta Rogal, Jarek Juraszek, Bernd Ensing, and Peter Bolhuis
J. Chem. Phys. 133, 174110 (2010)
27) The reweighted path ensemble.
Jutta Rogal, Wolfgang Lechner, Jarek Juraszek, Bernd Ensing, and Peter Bolhuis
J. Chem. Phys. 133, 174109 (2010)
26) Multiscale molecular dynamics and the reverse mapping problem.
Bernd Ensing and Steven O. Nielsen
Book chapter in Trends in Computational Nanomechanics (Challenges and Advances in Computational Chemistry and Physics, Volume 9) edited by T. Dumitrica (Springer, 2010), p. 25 – 60
25) Hydrolysis of cisplatin – a first-principles metadyamics study.
Justin Kai-Chi Lau and Bernd Ensing
Phys. Chem. Chem. Phys. 12, 10348-10355 (2010)
24) A simple coarse-grained model for self-assembling silk-like protein fibers.
Marieke Schor, Bernd Ensing, and Peter G. Bolhuis
Faraday Discuss. 144, 127-141 (2010)
2009
23) Toward a practical method for adaptive QM/MM simulations.
Rosa E. Bulo, Bernd Ensing, Jetze Sikkema, and Lucas Visscher
J. Chem. Theory Comput. 5, 2212 – 2221 (2009)
2007
22) Energy conservation in adaptive hybrid atomistic/coarse-grain molecular dynamics.
Bernd Ensing, Steven O. Nielsen, Preston B. Moore, Michael L. Klein, and Michele Parrinello
J. Chem. Theory Comput. 3, 1100 – 1105 (2009)
21) Proton shuttles and phosphatase activity in soluble epoxide hydrolase.
Marco De Vivo, Bernd Ensing, Matteo Dal Peraro, German A. Gomez, David W. Christianson, and Michael L. Klein
J. Am. Chem. Soc. 129, 387 – 394 (2007)
20) Coarse grained-to-atomistic mapping algorithm: a tool for multiscale simulations.
Steven O. Nielsen, Bernd Ensing, Preston B. Moore, and Michael L. Klein
Book chapter in Multiscale Simulation Methods for Nanomaterials, R. B. Ross and S. Mohanty, Editors. 2007, John Wiley & Sons, Inc., Hoboken, NJ, USA
2006
19) Self-assembling cyclic peptides: molecular dynamics studies of dimers in polar and nonpolar solvents.
Ekta Khurana, Steven O. Nielsen, Bernd Ensing, and Michael L. Klein
J. Phys. Chem. B 110, 18965 – 18972 (2006)
18) Formamide hydrolysis in alkaline aqueous solution: Insights from ab-initio metadynamics calculations.
Jochen Blumberger, Bernd Ensing, and Michael L. Klein
Communication in Angewandte Chemie Int. Edit. 45, 2893 – 2897 (2006)
17) Metadynamics as a tool for exploring the free energy landscape of chemical reactions.
Bernd Ensing, Marco De Vivo, Zhiwei Liu, Preston Moore, and Michael L. Klein
Acc. Chem. Res. 39, 73 – 81 (2006)
2005
16) Perspective on the reactions between F- and CH3CH2F: the free energy landscape of the E2 and SN2 reaction channels.
Bernd Ensing and Michael L. Klein
Cover article in Proc. Natl. Acad. Sci. USA 102, 6755 – 6759 (2005)
15) A recipe for the computation of the free energy barrier and lowest free energy path of concerted reactions.
Bernd Ensing, Alessandro Laio, Michele Parrinello and Michael L. Klein
J. Phys. Chem. B 109, 6676 – 6687 (2005)
14) Lipid bilayer perturbations around a transmembrane nanotube: a coarse grain molecular dynamics study.
Steve O. Nielsen, Bernd Ensing, Vanessa Ortiz, Preston B. Moore, and Michael L. Klein
Cover article in Biophys. J. 88, 3822 – 3828 (2005)
13) Structure and dynamics of model pore insertion into a membrane.
Carlos F. Lopez, Steve O. Nielsen, Bernd Ensing, Preston B. Moore, and Michael L. Klein
Cover article in Biophys. J. 88, 3083 – 3094 (2005)
2004
12) Computational study of phosphatase activity in soluble epoxide hydrolase: high efficiency through a water bridge mediated proton shuttle.
Marco De Vivo, Bernd Ensing, and Michael L. Klein
Communication in J. Am. Chem. Soc. 127, 11226 – 11227 (2004)
11) A minimum free energy reaction path for the E2 reaction between fluoro ethane and a fluoride ion.
Bernd Ensing, Alessandro Laio, Francesco L. Gervasio, Michele Parrinello, and Michael L. Klein
Communication in J. Am. Chem. Soc. 126, 9492 – 9493 (2004)
10) Methane to methanol oxidation by the hydrated iron(IV)oxo species in aqueous solution; a combined DFT and Car-Parrinello molecular dynamics study.
Bernd Ensing, Francesco Buda, Michiel C.M. Gribnau, and Evert Jan Baerends
J. Am. Chem. Soc. 126, 4355 – 4365 (2004)
2003
9) O-2 evolution in the Fenton reaction.
Francesco Buda, Bernd Ensing, Michiel C.M. Gribnau, and Evert Jan Baerends
Chem.Eur. J. 9, 3436 – 3444 (2003)
8) Fenton-like chemistry in water: Oxidation catalysis by Fe(III) and H2O2.
Bernd Ensing, Francesco Buda, and Evert Jan Baerends
J. Phys. Chem. A. 107, 5722 – 5731 (2003)
2002
7) Reaction path sampling of the reaction between iron(II) and hydrogen peroxide in aqueous solution.
Bernd Ensing and Evert Jan Baerends
J. Phys. Chem. A. 106, 7902 – 7910 (2002)
6) A Car-Parrinello study of the formation of oxidizing intermediates from Fentons reagent in aqueous solution.
Bernd Ensing, Francesco Buda, Peter E. Bloechl, and Evert Jan Baerends
Phys. Chem. Chem. Phys. 4, 3619 – 3627 (2002)
2001
5) Chemical involvement of solvent water molecules in elementary steps of the Fenton oxidation reaction.
Bernd Ensing, Francesco Buda, Peter E. Bloechl, and Evert Jan Baerends
Communication in Angewandte Chemie Int. Edit. 40, 2893 – 2895 (2001)
4) DFT study of the active intermediate in the Fenton reaction.
Francesco Buda, Bernd Ensing, Michiel C.M. Gribnau, and Evert Jan Baerends
Chem. Eur. J. 7, 2775 – 2783 (2001)
3) Solvation effects on the SN2 reaction between CH3Cl and Cl- in water.
Bernd Ensing, Evert Jan Meijer, Peter E. Bloechl, and Evert Jan Baerends
J. Phys. Chem. A 105, 3300 – 3310 (2001)
2000
2) Comparison of the accurate Kohn-Sham solution with the generalized gradient approximations (GGAs) for the SN2 reaction F- + CH3F -> FCH3 + F-: A qualitative rule to predict success or failure of GGAs.
Oleg V. Gritsenko, Bernd Ensing, Pieter R.T. Schipper, and Evert Jan Baerends
J. Phys. Chem. A 104, 8558 – 8565 (2000)
1998
1) A combined molecular dynamics-ab initio study of hydrogen molecule adsorption on ideal, relaxed and temperature-reconstructed MgO (111) surfaces.
Micael Baudin, Kersti Hermansson, Bernd Ensing, Maria Alfredsson, and Mark Wojcik
J. Chem. Phys. 109, 7515 – 7521 (1998)
Student research projects
If you are a UvA/VU bachelor or master student and you are interested in doing your research project in our group, please feel free to contact us for possibilities. We have almost always student projects ready related to the ongoing research lines in the group (see for examples the Research Interests above). If you like to work on machine learning and/or simulation method development, then a basic programming knowledge is definitely a pre. If you like to work on DFT-MD simulations of catalysis, electrochemistry, or enzymatic chemistry, then a good understanding of quantum chemistry is needed. In general, projects involving (classical) MD require an adequate grasp of thermodynamics and/or statistical mechanics.
Contact any of the group members:
- Ilaria Coratella
- Jacobus Dijkman
- Bernd Ensing
- Ferry Hooft
- Anastasia Kryachkova
- Matilde Mercuri
- Marta Monti
- Abhay Nagaraj
- Alec Noppe
Examples of available (master) research projects:
- Gen-AI for synthetic fuel catalysis
- AI-guided catalytic plastic liquefaction to produce sustainable molecules
Group photos






In the news
- February 2024
“AI helps chemists develop new, sustainable materials“, article on the FNWI News site on the startup of the Artificial Intelligence for Sustainable Molecules and Materials (AI4SMM) initiative, written by Harm Ikink.
- February 2024
“University Board visits Van ‘t Hoff Institute for Molecular Sciences“, article on the FNWI News site on our presentation of the AI4Science Initiative to the visiting Executive Board of the University of Amsterdam.
- January 2024
“AI marks a significant turning point in materials development“, commentary that I wrote for C2W Magazine on the revolutionary AI algorithms developed for material development by Google Deepmind (GNoME) and Microsoft Research (MatterGen). A Dutch version of the article is found here: “AI betekent een niet te onderschatten kentering in materialenontwikkeling“.
- December 2023
“Generatieve AI pakt rol in de duurzaamheidstransitie“, Interview and Sustainability Report (in Dutch) by Julia Krauwer Cas Bogaard for ABN AMRO.
- November 2023
“Four projects kick-start the Artificial Intelligence for Sustainable Molecules and Materials research programme“, highlight on the FNWI Newsletters about our four granted postdoctoral fellowships within the AI for Sustainable Molecules and Materials (AI4SMM) initiative.
- November 2023
“Companies and talent meet at ChemAI event“, based on an interview with Esther Thole for C2W Magazine about our ChemAI Event. For a Dutch version, see “ChemAI event brengt bedrijven en talent bij elkaar“.
- July 2020
“Humane-AI, on the spot“, an interview about the new AI4Science Laboratory for the Humane-AI news letter.
- June 2020
“AI4Science Lab sets out to take scientific data analysis to the next level“, a news item on the HIMS website by Harm Ikink about the AI4Science Laboratory.
- June 2020
Een Nederlandse versie van het artikel van Harm Ikink op de HIMS website is getiteld: “AI4Science Lab brengt analyse van wetenschappelijke data naar hoger niveau”.
- July 2019
Highlights on Chemistry world website, Solubility simulations solve mystery of why PEG dissolves in water but Keck clips don’t, on the C2W website, Electronen delen is geen oplossing, and on the UvA news site, Oplosbaarheidsmysterie opgelost and Solubility mystery solved, about our article in Nature Communications about the origin of the extremely different solubilities of polyethers in water.
- March 2019
Highlights in Science Daily and in Phys.org about our discovery with Lou Madsen and co-workers of the double helix structure in a synthetic macromolecule. Also MRS Bulletin published a news article about our work.
- September 2018
Interview in New Scientist magazine with Joris Janssen on computer simulations of water (in Dutch) “Met computersimulaties kun je de natuurwetten een beetje manipuleren“, New Scientist, September 2018, pages 44-45.
- 12 March 2018
Highlight in Chemistry World of the Royal Society of Chemistry on our paper with the groups of Woutersen and Angell on the liquid-liquid phase transition in an aqueous solution [Science 359, 1127-1131 (2018)]. Other media reporting on our work are found here: University of Amsterdam (Dutch) and C2W news (Dutch).
- 8 March 2018
The shapes of water: New research details water’s mysterious phase transitions on Phys.org after our paper in [Science 359, 1127-1131 (2018)].
- 4 February 2018
Interview on the Dutch radio station “Salto Radio” (podcast in Dutch) about “The special properties of water” (Radio Swammerdam 2018).
- 13 November 2014
Highlight at Phys.org on our paper with the Woutersen group on the orientational order in liquids [Phys. Rev. Lett. 113, 207801 (2014)]. The Volkskrant (Dutch Newspaper) reported our findings on 15 November. Our work was also highlighted on the websites of FOM and HIMS (in Dutch).
- 10 November 2014
Interview in Chemical & Engineering News on “Computational Chemistry: Ab initio nanoreactor points to unexplored reaction mechanisms” (Nature Chem. 2014).
- 28 July 2013
Highlight in ChemViews Magazine on our paper about sp3-Carbon-Based Liquid Crystals
[Chem. Eur. J. 19 (2013), 11577]. Our computer simulations showed that syndiotactic polycarbene polymers self-assemble into triple-helix structures, which explains the surprising liquid-crystal behavior of this unique material.
Biography
Bernd Ensing is a computational chemist who works on molecular modeling and machine learning algorithms to model, understand, and, ultimately, predict material behavior and (bio-) molecular processes. He is a full professor of AI for Chemistry at the Van ‘t Hoff Institute for Molecular Sciences and director of the AI4Science Laboratory, both at the University of Amsterdam. After his chemical engineering degree at the Technical University of Twente, Ensing received in 2003 his PhD in theoretical chemistry in the group of Prof. Baerends at the VU University Amsterdam (UvA). Subsequently, he held postdoctoral positions in the groups of Prof. Klein at the University of Pennsylvania and Prof. Parrinello at the ETHZ, before launching his research group at the UvA. His current research interests include improving electro- and bio-mimetic catalysis for sustainable chemistry and design of new molecules and materials for applications in the energy and health sectors.
AI4Science Laboratory
The AI4Science Laboratory is an interdisciplinary research group working on artificial intelligence and machine learning applied to all sorts of scientific research. The group is located in the LAB42 building and is connected to the different institutes of the Faculty of Science. Take a look at the AI4Science activities at the AI4science website.
Software
Our group develops various algorithms and computational tools to carry out advanced molecular simulations and to analyse simulation results. When we feel that a software implementation can be useful to others (and has some minimal stability, user-friendliness, and documentation), we have the habit to make it freely available to the community via our GitHub website.
Jobs
There are currently no funded PhD student or postdoctoral fellow positions open in the Ensing group. For positions in the Amsterdam Computational Chemistry group with other group leaders, please have look our vacancies page or the general UvA vacancies page.
Unfortunately, we are no longer able to reply the unsolicited applications and requests for positions in our group that we receive by email. Please accept our apologies in advance for not answering your email in case you apply for a funded PhD or postdoc position or request our advice on your academic career possibilities. When we have a vacancy in the group, it will be posted here and on the UvA vacancies page, with a link for submission of applications (but never through email).
Contact
- Contact
- Science Park 904, Room C2.223 (address details)
- Orcid page
- 0000-0002-4913-3571
- Google Scholar
- scholar.google.nl/BerndEnsing
- Mastodon
- @mstdn.science/@BerndEnsing
- LinkIn
- www.linkedin.com/in/berndensing
- Github
- github.com/Ensing-Laboratory
