Protein misfolding and aggregation are associated with some of the most intractable neurodegenerative disorders. The details of this aggregation process are still largely unknown, and are at the focus of current biochemical and biophysical research. A recent, surprising discovery could make it possible to shed new light on the mechanism of amyloid formation. It is generally believed that H/D-isotope substitution has no effect on molecular structure, and only on chemical rates if proton transfer is involved. However, it was recently found that amyloid fibrilization of insulin ismuch slower in D2O than in normal water, and for some proteins, such as elastin-like polypeptides or globular proteins, even the structure is different in H2O and D2O. The differences are currently believed to be due to the H/D-isotope effect on hydrogen bonds. These results might be the beginning of a paradigm shift regarding the use of D2O in chemistry (e.g. in protein NMR). On the other hand, investigating the structural and dynamic differences of fibrillization in water and heavy water can be a useful strategy to gain a better understanding on the aggregation process and the effects of the solvent.
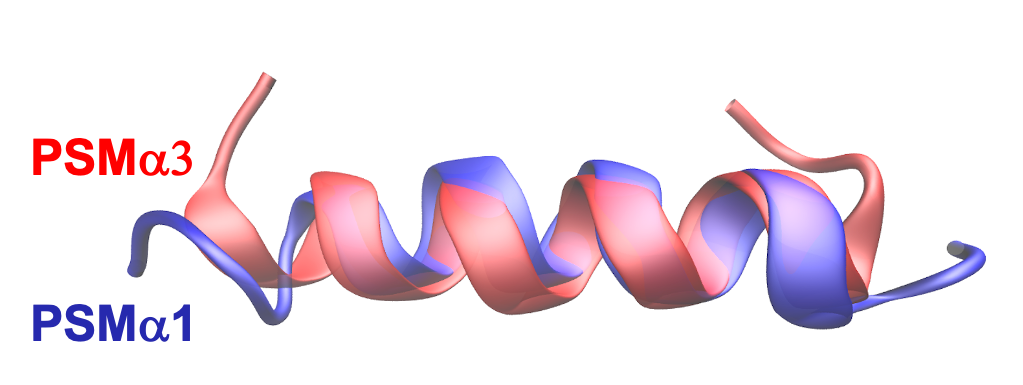
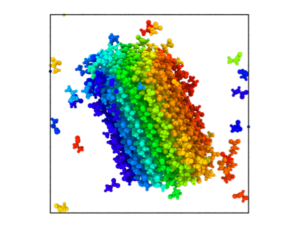
Goal: We aim to understand the aggregation dynamics of VEALYL, an amyloid-forming peptide derived from human insulin, in H2O and D2O by combining molecular dynamics (MD) simulations with experimental techniques, such as turbidity and fluorescence measurements, and linear and non-linear infrared spectroscopy. The experimental validation is done in collaboration with Dr. G. Giubertoni & Prof. Sander Woutersen.
Research questions
1. How does the solvent affect the thermodynamics and kinetics of aggregate formation & their structure?
2. What are the stabilizing interactions in aggregates in H2O and D2O?